Detection of genetic purging and predictive value of purging parameters estimated in pedigreed populations
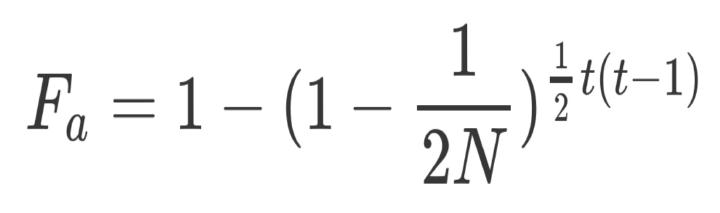
Abstract
The consequences of inbreeding for fitness are important in evolutionary and conservation biology, but can critically depend on genetic purging. However, estimating purging has proven elusive. Using PURGd software, we assess the performance of the Inbreeding-Purging (IP) model and of ancestral inbreeding ($F_{a}$) models to detect purging in simulated pedigreed populations, and to estimate parameters that allow reliably predicting the evolution of fitness under inbreeding. The power to detect purging in a single small population of size $N$ is low for both models during the first few generations of inbreeding ($t \approx N/2$), but increases for longer periods of slower inbreeding and is, on average, larger for the IP model. The ancestral inbreeding approach overestimates the rate of inbreeding depression during long inbreeding periods, and produces joint estimates of the effects of inbreeding and purging that lead to unreliable predictions for the evolution of fitness. The IP estimates of the rate of inbreeding depression become downwardly biased when obtained from long inbreeding processes. However, the effect of this bias is canceled out by a coupled downward bias in the estimate of the purging coefficient so that, unless the population is very small, the joint estimate of these two IP parameters yields good predictions of the evolution of mean fitness in populations of different sizes during periods of different lengths. Therefore, our results support the use of the IP model to detect inbreeding depression and purging, and to estimate reliable parameters for predictive purposes.